Introduction
Type 2 diabetes mellitus (T2DM) is the predominant form of diabetes worldwide and has been rapidly increasing in recent years. Currently, around 500 million people have diabetes, and this number is expected to increase by 25–51% between 2030 and 2045. These projections indicate that the incidence of diabetes will rise from 9.3% (463 million people) to 10.2% by 2030 and to 10.9% by 2045 (Saeedi et al., 2019). The onset of COVID-19 in early 2020 has highlighted the severe threat of complications and mortality associated with hyperglycemia in patients with T2DM (Holman et al., 2020; Zhu et al., 2020). As a result, there is an urgent need for the development of new therapeutics for T2DM.
The SIRT1 pathway is a commonly targeted signaling pathway in DM treatments due to its significant role in T2DM development. In mammals, Sirtuin 1 (SIRT1), a nicotinamide adenine dinucleotide (NAD(+))-dependent histone deacetylase (HDACs), modulates a wide range of cellular activities, including gene expression, DNA repair, metabolism, and oxidative response, and is thought to be a key regulator of longevity (Cohen et al., 2004 ; Baur et al., 2012). Dysregulation of SIRT1 has been implicated in the development of various human pathologies, including T2DM (Waldman et al., 2018). Interestingly, SIRT1 activity decreases with age, due to lowered concentration of NAD(+), whereas T2DM prevalence increases with age and peaks between 60 and 74 years old, likely due to insulin secretion/response abnormalities (Gong and Muzumdar, 2012). During the aging process, increased oxidative stress (OS) caused by reactive oxygen species (ROS) is accompanied by reduced cellular responsiveness to physiological stress. OS, combined with inflammation, contributes to the progression of insulin resistance in T2DM. Inflammatory mediators, such as IL-6, IL-8, TNF-, and MCP-1, are significantly elevated in the serum of aged individuals and contribute to insulin resistance and diabetes severity (Mei and Zheng, 2009; Olefsky and Glass, 2010). Interestingly, NFκB, considered the master regulator of proinflammatory cytokines, negatively correlates with SIRT1 in terms of its mechanism of action. Therefore, the discovery of novel compounds that target both SIRT1 and NFκB is a promising approach to identifying antiDM compounds associated with aging (Hung et al., 2015; Cuyàs et al., 2018; Fourny et al., 2019).
Curcuma, a genus in the family Zingiberaceae, is best known as a spice and food coloring agent in Asian cuisines, as well as a traditional medicine (Leong-Škornicková et al., 2015). Curcuma longa and Curcuma zedoaria are the most extensively studied among Curcuma species due to the high concentration of curcuminoids, including curcumin, desmethoxycurcumin, and bisdemethoxycurcumin (Itokawa et al., 2008). These curcuminoids are nontoxic and possess a broad spectrum of biological activities, including antiviral (Wahyuni et al., 2018), antibacterial, antioxidant (Kebede et al., 2021), and antihyperglycemic (Handajani and Narissi, 2015) effects. Curcuma zanthorrhiza Roxb is another important traditional herb in the Curcuma genus, besides C. longa and C. zedoaria. In Indonesia, C. zanthorrhiza is frequently consumed as “jamu”, a traditional health supplement, as many local societies believe it may maintain health or cure diseases (Shahid et al., 2021).
Similar to C. longa and C. zedoaria, C. zanthorrhiza displays antioxidant activities (Jantan et al., 2012), is nontoxic (Rahmayunita et al., 2018), and has been shown to possess antihyperglycemic and antiinflammatory activity (Kim et al., 2014). In addition, a previous study predicted that C. zanthorrhiza is involved in endogenous antioxidant action at the molecular level (Prasetyawan et al., 2022). C. zanthorrhiza contains a unique sesquiterpenoid, xanthorrhizol, which is absent in C. longa, and has beneficial effects on health (Jantan et al., 2012). However, most studies on the SIRT1/NFκB signaling pathway have focused on C. longa and curcumin as the main bioactive compound (Sahin et al., 2016; Zendedel et al., 2018). To our knowledge, no study has investigated how phytochemicals from C. zanthorrhiza may interact with the SIRT1/NFκB signaling pathway. Incorporating computer modeling in research has increased efficiency, decreased error rates, and avoided unnecessary time and material expenditures (Geris et al., 2016; Ononamadu and Ibrahim, 2021; Akhtar et al., 2022). Therefore, this study focuses on in silico approaches to discover C. zanthorrhiza bioactive compounds as SIRT1 activators and NFκB inhibitors and evaluate their toxicity.
Materials and methods
Ligand’s selection
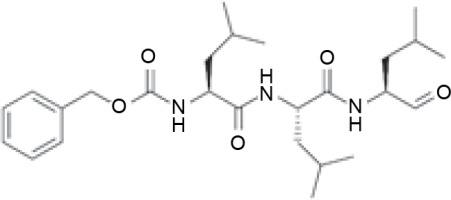
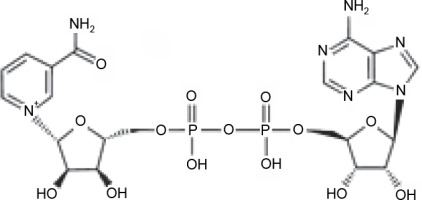
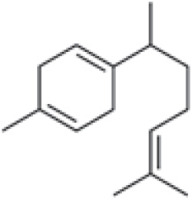
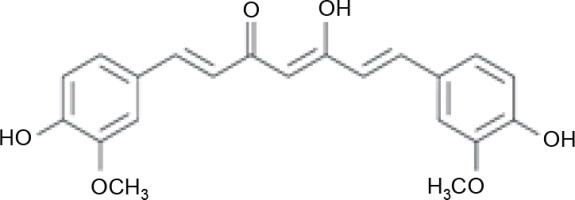
Three active compounds from C. zanthorrhiza, namely curcumin, beta-curcumene, and xanthorrhizol, were selected for molecular docking and are listed in Table 1. MG132, a molecule known to inhibit NFκB (Kadioglu et al., 2015), was chosen as the reference molecule. NAD(+) was selected as the reference molecule that activates SIRT1 (Azminah et al., 2019a). The 3D structures of the selected molecules were retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov/) in the sdf format. The downloaded molecules were converted into the pdb format using Pymol (Schr dinger Inc., LLC), and their energies were minimized using Open Babel in PyRx 0.8 (The Scripps Research Institute, California). Energy minimization is often performed before molecular docking to achieve stable conformation that is close to the original state in the biological system (Mackay et al., 1989).
Molecular docking simulation of SIRT1
The 3D structure of SIRT1 was obtained from the Protein Data Bank (PDB) (https://www.rcsb.org/). Water molecules were removed from the SIRT1 protein structures following standard procedures in PyMol. Three SIRT1 proteins, namely dimeric chains bound to NAD(+) (4I5I), heterodimeric chains (4ZZJ), and heterotrimeric chains (5BTR), were chosen for the present study, according to a previous report (Cuyàs et al., 2018). The grid position of 4I5I was set at x = 30.6373, y = –19.245, and z = 27.9433 with dimensions (Angstroms) x = 25.00, y = 25.2256, and z = 25.8823. The grid position of 4ZZJ was set at x = –0.8184, y = 45.1898, and z = –1.0706 with dimensions (Angstroms) x = 20.1723, y = 20.322, and z = 20.338. The grid position of 5BTR was set at x = –72.9063, y = 79.5449, and z = 15.3989 with dimensions (Angstroms) x = 20.4774, y = 19.6908, and z = 20.2907. The Discovery Studio v20 was used to evaluate the amino acids in SIRT1 that interact with curcumin, beta-curcumene, and xanthorrhizol. The protein-ligand interactions were visualized in Pymol.
Molecular docking simulation of NFκB
Three NFκB proteins were selected for the molecular docking simulation, namely NFκB p52–RelB complex (3DO7), NFκB p50–p65 heterodimer complexed to κB DNA (1VKX), and inhibitor of κB kinase beta (IκKB) (3RZF), based on a previous report (Kadioglu et al., 2015). The grid position of 3DO7 was set at x = 27.8426, y = 61.3128, and z = 75.5468 with dimensions (Angstroms) x = 20.2309, y = 22.2626, and z = 24.6053. The grid position of 1VKX was set at x = –1.9993, y = 34.3328, and z = 60.5516 with dimensions (Angstroms) x = 22.7518, y = 21.0639, and z = 20.2109. The grid position of 3RZF was set at x = 88.4660, y = –29.643, and z = 56.2548 with dimensions (Angstroms) x = 24.7078, y = 24.4270, and z = 24.5062. The NFκB proteins were collected, prepared, and visualized using the same procedures as SIRT1 described in the previous section.
Protein interaction and networking analysis
The STRING database (https://string-db.org/) was used to analyze protein–protein interactions (PPI) between SIRT1 and NFκB. In addition, the interaction of ligands with PPI was evaluated using STITCH (http://stitch.embl.de/).
Toxicity prediction
The three compounds from C. zanthorrhiza, tested for molecular docking, were evaluated for toxicity attributes, including LD50, class, hepatotoxicity, carcinogenicity, immunotoxicity, mutagenicity, and cytotoxicity using the Protox II database (https://tox-new.charite.de/protox_II/index.php?site=compound_input).
Results and discussion
Network analysis
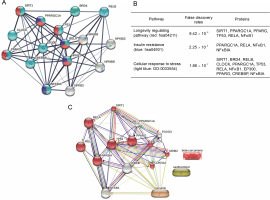
Protein–protein interaction analysis from the STRING database revealed several proteins involved in the SIRT1/NFκB pathways associated with longevity regulation, insulin resistance, and cellular response to stress (Fig. 2A). The three pathways involved in the SIRT1/ NFκB pathways covered three proteins, including peroxisome proliferator-activated receptor gamma coactivator 1-alpha (PPARGC1A), v-rel avian reticuloendotheliosis viral oncogene homolog A (RELA), and nuclear factor kappa b subunit 1 (NFKB1). On the other hand, SIRT1, PPARG, and TP53 were involved in two pathways related to longevity regulation and cellular response to stress (Fig. 2B). Further, STITCH analysis revealed nine proteins involved in the cellular response to stress in the SIRT/NFκB signaling pathway, including cyclin-dependent kinase inhibitor 1A (CDKN1A), murine double-minute 2 (MDM2), Forkhead box protein O1 (FOXO1), peroxisome proliferator-activated receptor gamma (PPARG), tumor protein p53 (TP53), nuclear factor-kappa-B-inhibitor alpha (NFKBIA), v-rel avian reticuloendotheliosis viral oncogene homolog B (RELB), RELA, and Sirtuin-1 (SIRT1) (Fig. 2C). Curcumin’s interaction in the SIRT1/NFκB signaling pathway was displayed with yellow lines, which means that curcumin could regulate the transcription of CDKN1A, MDM2, TP53 PPARG, and NFKBIA (Fig. 1C). However, betacurcumene and xanthorrhizol did not participate in the cellular response to stress pathways on the SIRT1/NFκB signaling axis, which displayed no lines connected between beta-curcumene and xanthorrhizol to the pathways built.
Fig. 1
Protein–protein interaction in SIRT1/NFκB signaling pathway; (A) SIRT1/NFKB axis generated by the STRING database in different pathways expected to interfere in T2DM; (B) Protein–protein is involved in the longevity regulating pathway, insulin resistance, and cellular response to stress; (C) Ligands and protein–protein interaction in the cellular response to stress (GO : 0033554) with false discovery rates 3.11×10-11; the lines color in Figure 1C are indicated: green – activation, red – inhibition, black – reaction, light blue – phenotype, navy – catalysis, pink – posttranslational modification, and yellow – transcriptional regulation
Docking simulation of selected C. zanthorrhiza bioactive compounds with SIRT1
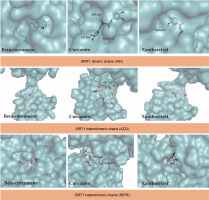
The docking results suggest that some bioactive compounds from C. zanthorrhiza have the potential to act as SIRT1 activators. Curcumin showed the strongest binding affinity following NAD(+), with binding energies of –7.9 kcal/mol to 4I5I, –5.1 kcal/mol to 4ZZJ, and –6.5 kcal/mol to 5BTR, followed by beta-curcumene and xanthorrhizol (Table 2).
Table 2
The binding affinities and amino acid interactions over molecular docking simulations of C. zanthorrhiza bioactive compounds against SIRT1
The molecular docking simulation results showed that beta-curcumene, curcumin, and xantorrizhol were located at the same binding site as NAD(+). Gly364, located on the dimeric chains of SIRT1, was found to be the amino acid involved in all ligand interactions. (Table 2). For the SIRT1 heterodimeric chains, both Asn226 and Glu230 were involved in all ligand interactions (Table 2). However, in the ligands-SIRT1 heterotrimeric complexes, Thr177 and Glu230 did not participate in the beta-curcumene and xanthorrhizol complexes, respectively (Table 2).
The results indicate that the bioactive compounds selected from C. zanthorrhiza show potential as candidates for activating SIRT1. The SIRT1 active site consists of two regions: the C-terminal region and N-terminus region. The C-terminal region contains 25 residues that are necessary for SIRT1 activity (ESA). These residues stabilize the catalytic core and enhance SIRT1 activity. The N-terminus region consists of a small domain with three helices that bind and activate SIRT-activating compounds (STACs). This region is called the STAC-binding domain (SBD), and it serves as a docking site for the compounds, positioning them at the active site entrance of the catalytic domain/substrate complex (Azminah et al., 2019b; Weiss et al., 2022). Additionally, Gly364 is an interactive site in SIRT1 that increases SIRT1 deacetylase activity, promoting its role in regulating metabolism and gene expression (Shen et al., 2019). Glu230 is crucial for stabilizing the deacetylase activity during SIRT1 activation, and substitution of Glu230 may disrupt enzymatic activity, decreasing the electrostatic interaction at the substrate-binding site (Azminah et al., 2019a). In line with our findings, another study reported that hyperoside and quercetin bind to Glu230 and Asn226 in SIRT heterodimeric chains (Rahayu et al., 2022). Additionally, a previous study highlighted the importance of Asn226 and Glu230 in stabilizing the allosteric site of SIRT1 (Azminah et al., 2019b).
A previous report suggested that curcumin increases 5′-adenosine monophosphate-activated protein kinase (AMPK) levels, followed by the elevation of superoxides and ATP levels to stimulate NAD(+) and SIRT1 activation (Zendedel et al., 2018). Another study reported that bioactive compounds from C. zanthorrhiza promoted SIRT1 expression in HEK293 cell lines (Zhang et al., 2015). Therefore, we hypothesize that the mechanism of action of curcumin is influenced by its structure. Curcumin has two aromatic ring systems with o-methoxy phenolic groups and seven carbon linkers containing an α, β-unsaturated diketone moiety. The presence of the β-diketone structure and hydroxyl groups in its aromatic ring is predicted to be an essential factor influencing curcumin's potential (Malik and Mukherjee, 2014; Priyadarsini, 2014). Furthermore, a previous study reported that curcumin may protect against OS by activating Nrf2, the master regulator of antioxidant enzymes (Suprihatin et al., 2017).
Xanthorrhizol is a sesquiterpene phenol on the bisabolene skeleton, which is predicted to influence its antioxidant properties (Oon et al., 2015). We presumed that the chemical structure might impact the docking result, which was confirmed by our results showing that curcumin had a stronger affinity, than xanthorrhizol, to SIRT1 (Table 2). Moreover, curcumin had the most negative binding affinity values, indicating that the interaction between curcumin as a ligand and SIRT1 dimeric chains (4I5I), SIRT1 heterodimeric chains (4ZZJ), and SIRT1 heterotrimeric chains (5BTR) as proteins will reach an equilibrium state eventually if the binding affinity values or Gibb's energy tends to have a negative value (Atho’illah et al., 2021; Pertami et al., 2021).
Docking simulations of selected C. zanthorrhiza bioactive compounds with NFκB
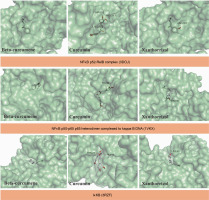
In this study, the selected compounds of C. zanthorrhiza showed potential as NFκB inhibitors. Curcumin demonstrated the strongest binding affinity to p52/RelB (–6.5 kcal/mol) and p50–p65 NFκB (–6.4 kcal/mol) after MG-132, followed by beta-curcumene and xanthorrhizol. Furthermore, xanthorrhizol showed a stronger binding affinity (–7.5 kcal/mol) than beta-curcumene and curcumin with IκK complexes (Table 3).
Table 3
The binding affinity calculations and amino acid interaction over molecular docking simulations of C. zanthorrhiza bioactive compounds against NFκB
Molecular docking simulations demonstrated that beta-curcumene, curcumin, and xanthorrhizol docked at the same binding site as MG-132. Asp219, Ser188, and Ser222 were involved in all ligand interactions with p52 RelB NFκB complexes, except for MG-132 (Table 3). Curcumin and xanthorrhizol interacted with the same amino acid in the p50–p65 NFκB complexes, Pro189 (Table 3). Asp166, Glu149, and Gly22 from IκK were interacting with all ligands except MG-132 (Table 3).
The current study demonstrates the potential of C. zanthorrhiza bioactive compounds as NFκB inhibitor candidates. It has been shown that the selected C. zanthorrhiza bioactive compounds interact with the residues at pharmacophore regions of the p52–RelB complex (Kadioglu et al., 2015). For example, beta-curcumene interacted with Arg54 and Glu58 from p52. Curcumin interacted with Tyr55 from p52 and Arg333 from RelB, while xanthorrhizol interacted with Arg54 and Glu58 from p52. In addition, Arg54 and Glu58 were found to be involved in the base-specific polar interaction with the core sequence flanking the CCC:GGG, while Tyr55 was responsible for contacting the hydrophobic carbons and rings of DNA bases, thus mediating sequence-specific van der Waals interactions (Abdelhameed et al., 2021). Among the amino acids, Ser222 was most frequently involved in the interaction with beta-curcumene–p52 complexes, curcumin–p52 complexes, and xanthorrhizol–p52 complexes. Ser222 is the interdomain linker that makes water-mediated contacts with the DNA backbone (Cramer et al., 1997).
In the p50–p65 heterodimer complex, beta-curcumene interacted with Gln247 from p65, while curcumin interacted with Arg33 and Tyr36 from p65. Tyr36 is essential for base-specific contact for p65 polyubiquitination, and its substitution with Tyr36 diminished p65 binding to the canonical κB sites, in both homodimers and heterodimers with p50 (Saccani et al., 2004). Moreover, it is one of the essential residues in the p50–p65 heterodimer. Arg33 of the RelA chain facilitated the recognition of guanine (gua)25′ and gua24′ in both the Core II and Core I regions (Stroud et al., 2009). On the other hand, Gly24, Ile165, and Asp166 of IκK proteins may be important in the ATP binding sites. Asp residues formed a polar contact with ATP-phosphatase directly or through magnesium atoms in the catalytic loop (Sala et al., 2011). Thus, the interaction between C. zanthorrhiza bioactive compounds and key NFκB residues was deemed adequate to prevent NFκB recognition on the κB DNA sites.
Oral toxicity prediction of C. zanthorriza bioactive compounds
The selected C. zanthorrhiza bioactive compounds were predicted to have oral toxicity of class 4 (curcumin and xanthorrhizol) and class 5 (beta-curcumene) (Table 4). Class 4 compounds are harmful if swallowed, with an LD50 ranging between 300 and 2000 mg/kg, while class 5 compounds may be harmful if swallowed, with an LD50 ranging between 2000 and 5000 (Banerjee et al., 2018). Surprisingly, curcumin was predicted to have one toxicity attribute, indicating immunotoxicity, while betacurcumene and xanthorrhizol did not show any toxicity features (Table 4).
Table 4
Oral toxicity prediction using Protox II
| Compounds | LD50 [mg/kg] | Class | Hep | Cg | Im | Mtg | Cy |
|---|---|---|---|---|---|---|---|
| Beta-curcumene | 3650 | 5 | (–) | (–) | (–) | (–) | (–) |
| Curcumin | 2000 | 4 | (–) | (–) | (+) | (–) | (–) |
| Xanthorrhizol | 2000 | 4 | (–) | (–) | (–) | (–) | (–) |
[i] Note: Hep – hepatotoxicity, Cg – carcinogenicity, Im – immunotoxicity, Mtg – mutagenicity, Cy – cytotoxicity, (+) – active, (–) – inactive; Class 1 – fatal if swallowed (LD50 ≤ 5), class 2 – fatal if swallowed (5<LD50 ≤ 50), class 3 – toxic if swallowed (50<LD50 ≤ 300), class 4 – harmful if swallowed (300<LD50 ≤ 2000), class 5 – may be harmful if swallowed (2000<LD50 ≤ 5000), and class 6 – nontoxic (LF50 > 5000)
It has been shown that curcumin uptake of up to 8 g per day is safe for humans for three months (Zheng and McClements, 2020). Moreover, no mortality was observed after xanthorrhizol treatment at a dose of 500 mg/kg in mice, and it was estimated that it is still safe to consume up to 619 mg/kg of xanthorrhizol (1 mg of C. zanthirrhiza ethanolic extract contains around 0.1238 mg of xanthorrhizol) in 5 g/kg of C. zanthorrhiza extract (Oon et al., 2015). There is a lack of information regarding the toxicity and safety of beta-curcumene (Dosoky and Setzer, 2018). However, the toxicity prediction conducted in this study also proves that the bioactive compounds from C. zanthorrhiza are considered safe to consume.
Conclusions
The present study has demonstrated the importance of C. zanthorrhiza bioactive compounds such as beta-curcumene, curcumin, and xanthorrhizol as promising candidates to reduce the negative impact of DM associated with aging via the SIRT1/NFκB signaling pathway through computational modelling. Curcumin was found to have a stronger binding affinity to SIRT1 (4I5I, 4ZZJ, and 5BTR) and NFκB (3DO7 and 1VKX) than betacurcumene or xanthorrhizol. However, Xanthorrhizol was found to be a stronger inhibitor of IκK (3RZF) than beta-curcumene or curcumin. Based on toxicity prediction, beta-curcumene, curcumin, and xanthorrhizol belong to classes 4 or 5, indicating low toxicity. The present study’s results suggest that curcumin may be a potent SIRT1 activator and NFκB inhibitor. However, herbal interactions with other proteins through different signaling pathways, pharmacokinetic/pharmacodynamic studies of C. zanthorrhiza bioactive compounds such as beta-curcumene, curcumin, and xanthorrhizol, and other animal or clinical studies require further investigation.